

Document Type : Review
Authors
1 Urology Research Center, Tehran University of Medical Sciences, Tehran, Iran
2 Department of Pathology, Isfahan Kidney Diseases Research Center, Isfahan University of Medical Sciences, Isfahan, Iran
3 Urology and Nephrology Research Center, Hamadan University of Medical Science, Hamadan, Iran
4 Department of Geriatric Medicine, School of Medicine, Tehran University of Medical Sciences, Tehran, Iran
5 Medicine Department, Isfahan University of Medical Sciences, Isfahan, Iran
Abstract
Highlights
Keywords
Introduction
It is well known that microbes and humans have closely related and healthy bodies have a variety of microorganisms (1, 2). Microbes play a significant role in human health and disease and can cause bodily homeostasis or activities such as metabolism and defense (3). Today, the terms “microbiome” and “microbiota” are widely used in clinical science. Microbiota refers to the collection of microorganisms that reside in the part of the body, while microbiome means the pool of genomes and genes of the organisms that live in the host (4).
In 2008, the National Institute of Health conducted the human microbiome project (HMP) to study the human microbiome in 242 healthy individuals (5). In this interdisciplinary project, the microbiome of different sites of the body (including skin, gastrointestinal (GI) tract, oral cavity, nasal cavity, and vagina) was examined, and its relationship with human health and disease was analyzed (5). Initially, urine and urinary tract were not included in the HMP studies. One of the reasons for this was that urinary tracts were maintained sterile, and urinary disorders were thought to be caused by the invasion of an external pathogen (6).
For many years, microbiology was based on the technique of bacterial culture. Accordingly, the presence of bacteria was confirmed if the culture was positive. Currently, with the advent of methods beyond culture, the presence of bacteria in various parts of the body has been identified even under health conditions. Methods such as metagenomic analysis of bacteria have led to the discovery of complex or slow-growing bacteria in areas considered sterile (7-9).
Microbiota and the Urinary Tract
Advances in quantitative urine culture (EQUC), 16S rRNA, and metagenomic sequencing have increased knowledge of the specific microbiota of urine (9-12). In such studies, urine is taken directly from the bladder (with direct suprapubic aspiration) so that the examined microbiota is exclusively related to the urine and is not contaminated (9). The EQUC technique helps to analyze live bacteria not identified by traditional cultures (12). Because this culture technique uses a large amount of urine in specific culture media and temperatures (13). Also, the 16S rRNA gene is highly conserved and is a good target for bacterial sequencing. Because the presence of 9 hypervariable gene regions (V1-V9) in this conserved gene makes it possible to classify the bacteria in the sample by sequencing even one region of these genes (14, 15). The stages of urinary microbiota analysis are shown schematically in Figure 1. Therefore, with the help of these methods, live hard/slow bacteria could be evaluated in the urine, and the microbiota pattern of urine and urinary tract were determined (12). This microbiota pattern plays an important role in urinary tract health and disease (8, 16, 17). The association of microbiota with urinary tract health/disease is directly affected by microbiota changes in a variety of conditions, including seasonal, geographical, and nutritional conditions (3). Therefore, more efficient diagnosis and treatment can be achieved by targeting specific microorganisms present in the urinary system and their environmental conditions.
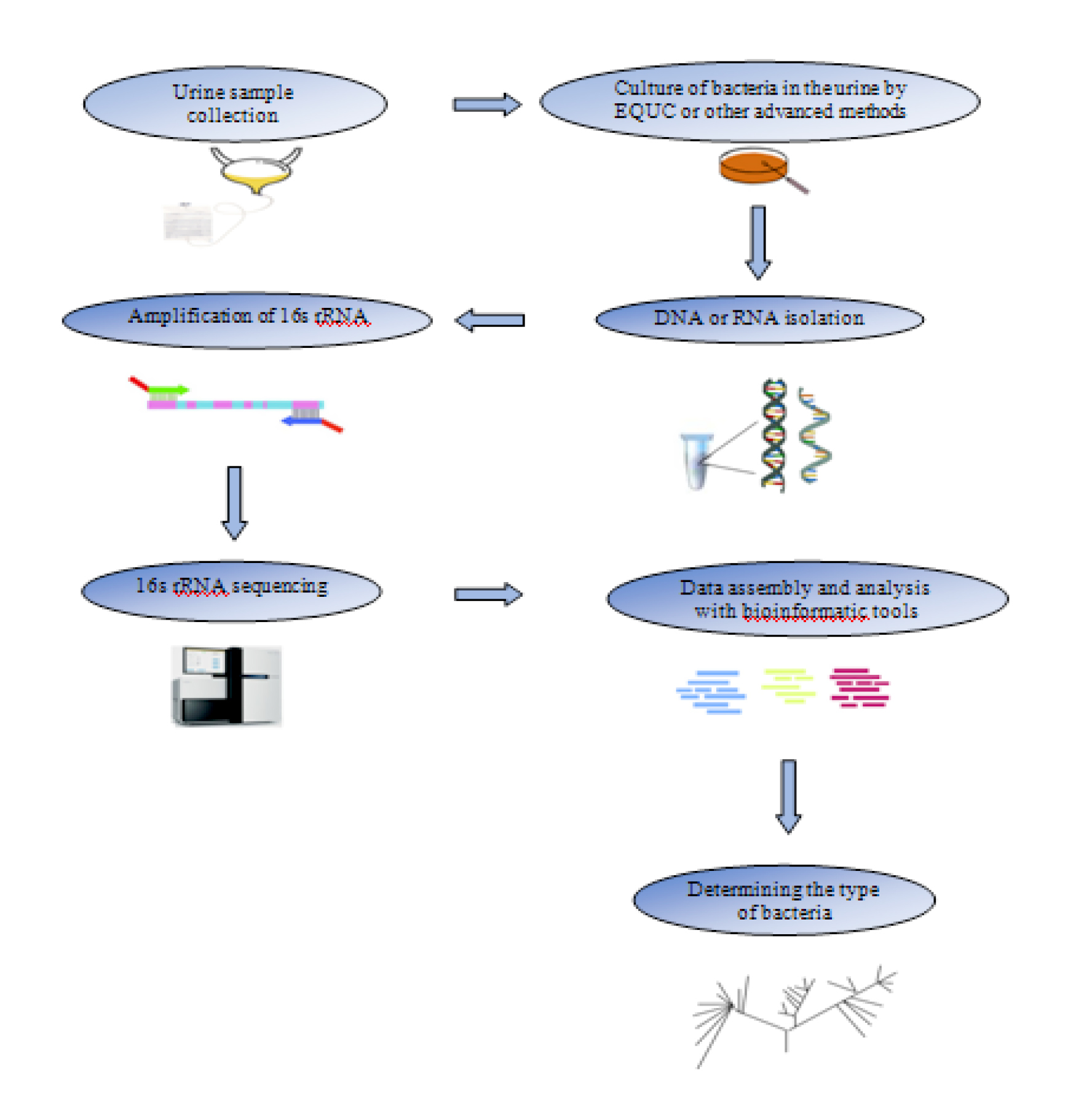
Figure 1. Schematic steps of the urine microbiota examination
Urinary Microbiota and the Overactive Bladder
Recent studies have shown that alterations in the urinary microbiota can be associated with many urological diseases, such as overactive bladder, interstitial cystitis, chronic prostatitis, and bladder cancer (14, 16, 18).
The international continence society (ICS) defines overactive bladder (OAB) as the sudden urge to urinate, with or without urge incontinence, often accompanied by frequency and nocturia in the absence of identifiable causes such as urinary tract infection (19, 20). OAB has been seen in both sexes, and its prevalence increases with aging (21).
In addition to the financial burden, OAB has an adverse effect on people's quality of life, sexual function, sleep, and physical and social activities. Also, it affects people's mental health due to disappointment, anxiety, shame, and stress (22, 23). OAB diagnosis is not a standard procedure and is performed only by physical examination and voiding diary. However, the possibility of infection and abnormal urinary tract pathology should be excluded (14).
The etiology and effective treatment of OAB is not yet fully clear. Possible causes are neuromuscular and muscarinic receptor dysfunction, but given that in many OAB patients, anti-muscarinic receptor, Botox, or other treatments are ineffective. Therefore, OAB can be caused by other reasons or a complex of reasons, and microbiota can be one of them (17, 24, 25). The articles related to the relationship between microbiota and active bladder, which are reviewed in this study, are summarized in Table 1.
Table 1. Summary of publications, addressing the role of the urinary microbiota in OAB patients
|
References |
Sample size |
Country |
Sample collection |
Analysis technique |
Main finding
|
|
Hilt et al., 2014 (24) |
41 OAB women and 24 healthy women |
USA |
Transurethral catheterization |
EQUC1 and 16S rRNA sequencing |
Common genera: Lactobacillus, Corynebacterium, Streptococcus, Actinomyces, Staphylococcus, Aerococcus, Gardnerella, Bifidobacterium, Actinobaculum Unique genera in the OAB group: Aerococcus and Actinobaculum |
|
Siddiqui et al., 2014 (26) |
Two samples from one OAB woman |
Norway |
Not described |
16S rDNA sequencing |
Streptococcus, Atopobium, Ureaplasma, Prevotella, Bacteroides |
|
Wu et al., 2017 (27) |
30 OAB women and 25 healthy women |
China |
Transurethral catheterization |
16S rRNA sequencing |
Decreased genera in the OAB group: Prevotella, Dialister, Fusobacterium, Jonquetella, Campylobacter, Finegoldia, Anaerococcus, Lactobacillu, Pyramidobacter, Ureaplasma, Enterococcus, Novosphingobium, and Lactococcus Increased genera in the OAB group: Sneathia, Staphylococcus, Proteus, Helcococcus, Gemella, Mycoplasma, and Aerococcus |
|
Gill et al., 2018 (28) |
24 OAB women and 22 healthy women |
UK |
MSU2 collection |
Urothelial-cell-sediment culture |
Common bacteria in the OAB group: E. coli, Enterococcus faecalis, Proteus, Klebsiella, Enterobacter, Serratia and Pseudomonas Common bacteria in health group: Staphylococcus, Streptococcus, Citrobacter, and Lactobacillus |
|
Perovic et al., 2019 (29) |
6 OAB women |
Portugal |
MSU2 collection |
Optimized CULT and 16s rRNA sequencing
|
Common bacteria: Bifidobacterium species and Corynebacterium aurimucosum Common bacteria in severe OAB patients: Gardnerella vaginalis and Ureaplasma urealyticum |
EOUC: Enhanced Quantitative Urine Culture; MSU: Midstream Specimen of Urine; OAB: Over Active Bladder
Hilt and colleagues used 16S rRNA gene sequencing and an EQUC protocol to examine live bacteria in the urine of 41 women with OAB and 24 healthy women. According to the EQUC technique, there were 52 positive culture samples (80%), while 92% were negatively reported by the standard culture protocol. Among the isolated genera, Lactobacillus (15%) was the most prevalent, followed by Corynebacterium (14.2%), Streptococcus (11.9%), Actinomyces (6.9%), and Staphylococcus (6.9%). In patients with OAB, Aerococcus and Actinobaculum were unique genera that were not seen in normal samples. The Hilt study showed that urine contains live bacteria originating from its microbiota and can be amazing in OAB (24).
In 2014, another study was performed on a 61-year-old woman with urinary symptoms and OAB for many years. In the first stage, bacteriuria (Streptococcus) was diagnosed with standard bacterial cultures and treated with antibiotics. By performing culture-independent 16S rDNA sequencing on the woman's urine, a set of different bacteria, including fastidious bacteria, were identified. A year later, the patient was re-examined for severe urinary symptoms, and no infection was reported by standard culture methods. But, the 16S rDNA results showed the same pattern as last year. Due to negative urine culture, the persistence of urinary symptoms, and the similarity of 16S rDNA results with the previous year, the association between microbiota and OAB becomes stronger (26).
In 2017, Wu and colleagues studied urinary microbiota in people with OAB. This study analyzed the urine of 30 patients with OAB and 25 healthy individuals (none of them had a urinary tract infection). 16S rRNA Sequencing results showed that 13 genera, including Prevotella, Dialister, Fusobacterium, Jonquetella, Campylobacter, Finegoldia, Anaerococcus, Lactobacillus, Pyramidobacter, Ureaplasma, Enterococcus, Novosphingobium, and Lactococcus were decreased in the OAB patients compared to the control group. In contrast, seven genera, including Sneathia, Staphylococcus, Proteus, Helcococcus, Gemella, Mycoplasma, and Aerococcus, in the OAB group were increased compared to the control group. Hence, there was a significant difference between the microbiota of the OAB group and the control. This study showed that certain species, such as Aerococcus, were associated with some specific urinary symptoms. Also, the urinary microbiota in OAB people is less diverse and is poorer in lactobacilli than in normal people. Finally, Wu et al., concluded that Lactobacillus's high diversity and dominance in microbiota were two protective factors against OAB (27).
In another study, the control group evaluated urinary tract microbiology activity for 12 months in OAB patients. Microbial evaluation in this study was performed using the culture of urothelial cells from the urine sediment, and its results were compared with the standard culture method. Among patients with OAB, E. coli was the most common species, followed by Enterococcus faecalis, Proteus, Klebsiella, Enterobacter, Serratia, and Pseudomonas. On the other hand, the most common species in the control group were Staphylococcus, Streptococcus, Citrobacter, and Lactobacillus. In this study, there are recurring and consistent differences in microbial load and dispersion of isolated species in the OAB and control groups. The difference in the amount and type of bacterial colonization in the patients and healthy individuals indicates the effect of urinary tract microbiota on the incidence and severity of the disease (28).
In 2019, Gill et al. examined urinary microbiota in six OAB patients without urinary tract infections using optimized CULT and 16S rRNA sequencing. These methods detected a huge number of bacteria in all patients with no criteria for urinary tract infection, which was related to 36 bacterial genera. Most of these bacteria are related to urinary tract infections, while standard culture methods have been unable to detect them. The most common bacteria in this study were Bifidobacterium species (28%) and Corynebacterium aurimucosum (22%). In patients with severe symptoms of OAB, Gardnerella vaginalis (46%) and Ureaplasma urealyticum (44%) were more common. Finally, Gill et al., conclude that urinary microbiota analysis is required to diagnose and treat OAB using new culture and sequencing methods (29).
Conclusions
Today, it has been proven that urine and the urinary tract are not sterile and have unique microbiota. These studies provide strong evidence that urinary microbiota changes in OAB patients. Therefore, the urinary microbiota was associated with OAB and may be used as a diagnostic and therapeutic marker. Studies with a large sample size can lead to a microbiota signature in OAB disease that can distinguish between the disease and other urinary tract diseases with overlapping symptoms.
Authors' contributions
All authors contributed equally.
Acknowledgments
Special thanks to Urology Research Center (URC), Tehran University of Medical Sciences, Tehran, Iran.
Conflict of interest
All authors claim no competing interest in this surgery case report.
Funding
There was no funding.
Ethics statement
Not applicable.
Abbreviations
EQUC Quantitative urine culture
GI Gastrointestinal
HMP Human microbiome project
ICS International continence society
OAB Overactive bladder