

Document Type : Review
Authors
1 Urology Research Center, Tehran University of Medical Sciences, Tehran, Iran
2 Isfahan Kidney Diseases Research Center, Isfahan University of Medical Sciences, Isfahan, Iran.
3 UroScience and Department of Surgery (Urology), School of Medical Sciences, University of Campinas, Unicamp, and Pontifical Catholic University of Campinas, PUC-Campinas, Campinas, São Paulo, Brazil
Abstract
Highlights
Keywords
Main Subjects
Introduction
Apoptosis plays a significant role in developing the pathogenesis of various cancers, autoimmune diseases, and viral infections. Besides, Autophagy is the main intracellular path for the damage and recycling of long-lived proteins and damaged organelles. In late 2019, a new coronavirus was recognized as causing a group of pneumonia cases in Wuhan, China. The disease spread rapidly, resulting in an epidemic in China and reports of worldwide outbreaks. In February 2020, the World Health Organization identified COVID-19, which stands for Coronavirus 2019. Studies on COVID-19 and producing a vaccine are in progress. This review attempts to summarize the present finding, focusing on the mechanisms and function of the coronavirus protein as a basis for better understanding the virus and vaccine production.
Methods
This study is a comprehensive review of findings on coronavirus. We summarized the available information on aspects of virology, mechanism, immune response, hormonal immunity, apoptosis and autophagy, and vaccine production. The issue is discussed on theoretical knowledge, practical aspects, and new findings to the date of the review (December 2020).
Virology
Coronaviruses have unique properties, including enveloped, non-segmented, and positive-sense single-stranded viruses. The genome size of the coronavirus is between 26 and 32 kb, the largest genome of RNA viruses. The replicase gene encodes the key purpose of coronavirus RNA production.
The coronavirus genome has 14 open reading frames (ORFs) that encode 27 proteins. The orf1ab and orf1a genes positioned at the '5' end of the genome; encode the pp1ab and pp1a proteins, respectively. Together, these two genes express 15 non-structural proteins, the nsp12-nsp16 and nsp1-nsp10. On the other hand, at the '3' end of the genome, four structural proteins, including spike surface protein (S), coat protein (E), membrane protein (M), nucleocapsid protein (N), and eight subunit proteins are coded including 3a, 3b, p6, 7a, 7b, 8b, 9b and orf14 (Figure1 and 2). The new coronavirus genome in some regions is significantly different from the SARS. The gene is composed of more than 20000 nucleotides. The unique size of the genome and the mechanism of particularly long-genomic RNA production in coronavirus are related to the production of non-structural proteins (protein 15 (nsp15)) that are unprecedented among RNA viruses. Many of these replicate fission products themselves are multipart proteins that enhance the functional and interactional complexity of the protein. Structural studies have shown that naps form dimers after being released from precursor molecules.
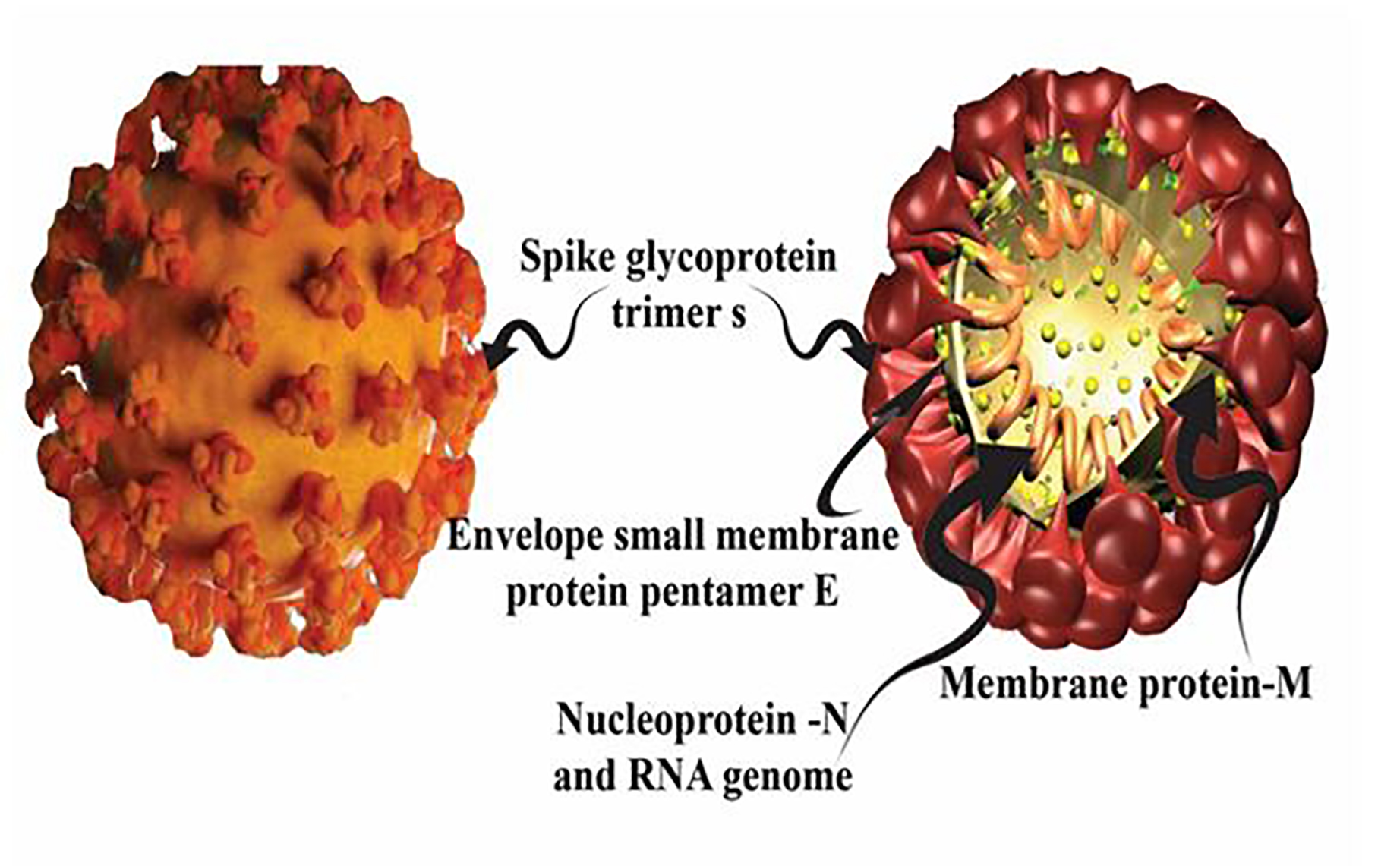
Figure 1. Coronavirus proteins. Schematic diagram of structural proteins of coronavirus that are encoded by structural genes, 1- S: spike glycoprotein trimer s, 2- M: membrane protein, 3- E: envelope small membrane protein pentamer, and 4- N: nucleocapsid protein or Nucleoprotein-N and RNA genome
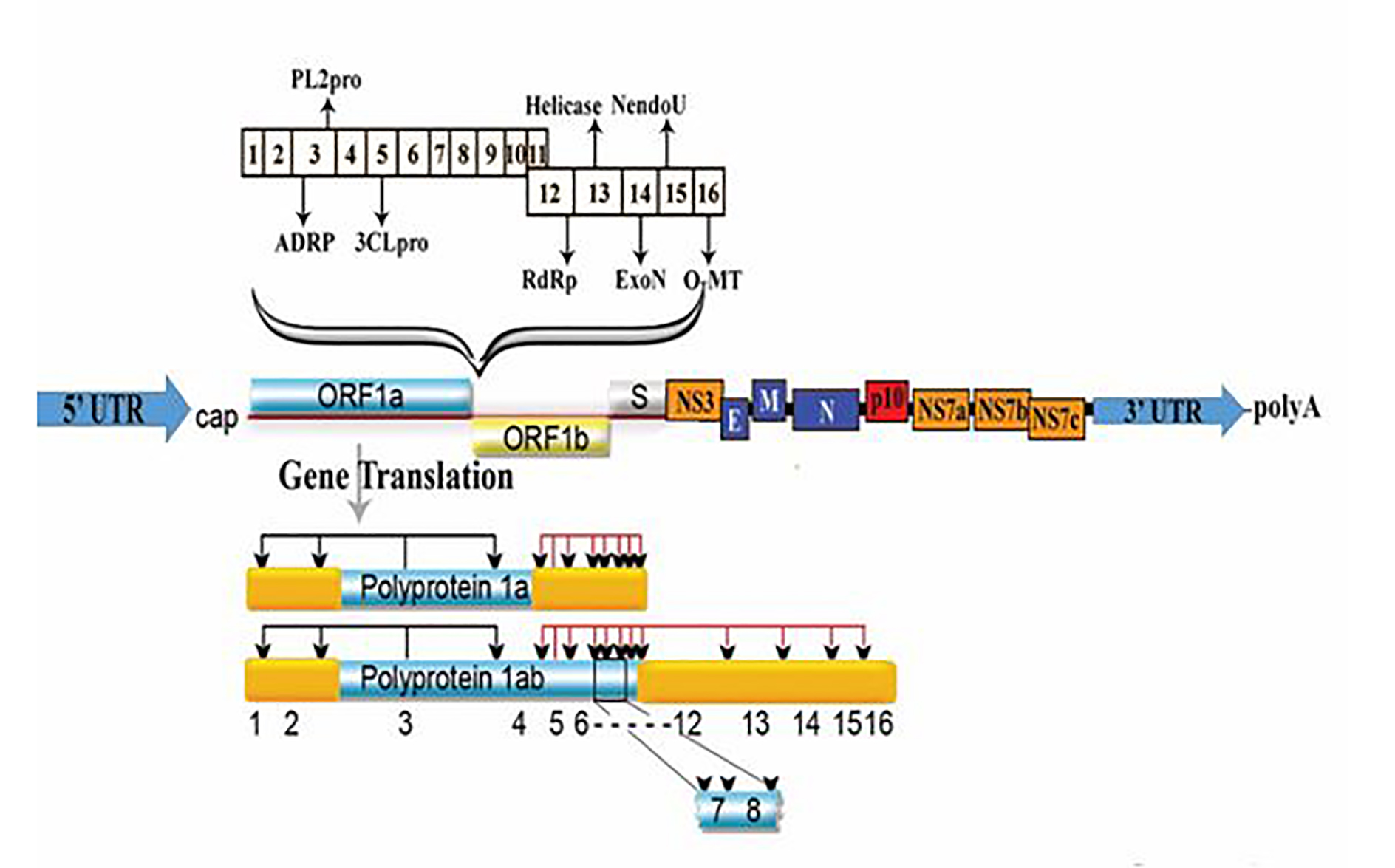
Figure 2. Coronavirus genome. The positive-sense single-stranded RNA genome of Covid-19 encodes two very large genes, ORF1a and ORF1b (as leader sequence encoding ingredients of the replication and or transcription assembled). It then encodes sixteen non-structural proteins that are extremely protected across all coronaviruses. There are eight subsidiary proteins containing 3a, 3b, 6, 7a, 7b, 8a, 8b, and 9b. a 5’ cap, leader sequence, UTR, 3’ UTR, genes for structural and accessory protein, a replicase gene and a poly-A tail modified by Coronavirus genomic RNA.
The pp1a/pp1ab precursors and processing products become large membrane-bound complexes that accelerate the reactions involved in RNA production and transcription and appear to have other functions in the virus life cycle. Coronaviruses also show alteration of the ribosomal framework and polymerase inhibition as part of a complex transcriptional cycle. The coronavirus has a nucleotide capsule containing genomic RNA and phosphorylated nucleocapsid proteins encapsulated in a bilayer phospholipid (1). These viruses have two different surface proteins named after the same appearance. The family of coronaviruses is genotyped and serologically divided into three types: coronavirina and torvirina. Coronavirinae is classified into three subsets: alpha, beta(isolated from mammalian hosts, including humans), gamma (isolated from avian hosts) (2), and delta. This classification of coronaviruses is based on antigenic relationships and comparing their genomic sequences. Common types of coronaviruses include alpha coronaviruses (229E, NL63) and beta coronaviruses (OC43, SARS, MERS, and HKU1). Complete genome sequencing and phylogenetic analysis showed that the coronavirus that causes COVID-19 is a type of beta coronavirus in the same subgroup of acute respiratory syndrome virus (as in several other bat coronaviruses), but takes in a different group (Figure 3). The Middle East Respiratory Syndrome, another beta coronavirus, is far more associated with the new virus. The RNA sequence closely resembles the two-corona virus of the bat, and the bats appear to be the main source. It is unclear whether the COVID-19 virus is transmitted directly from bats or via another mechanism (e.g. through the host).

Figure 3. Coronavirus phylogenetic tree. The scientific name chosen for coronaviruses is Orthocoronavirinae or Coronavirinae. Coronaviruses are from the Coronaviridae family, order Nidovirales, and realm Riboviria. The Nidovirales order has four families: Arteriviridae, Roniviridae, Mesoniviridae, and Coronaviridae. Coronaviridae itself includes two subfamilies: Torovirinae and Coronavirinae. Coronavirinae is divided into alpha and beta, which can infect mammals, including humans, and gamma and delta, which can infect birds. Genus: Alpha coronavirus: Species: HCoV-229E, HCoV-NL63, Mi-BatCoV-1A, Mi-BatCoV-HKU8, PEDV, Rh-BatCoV-HKU2, Sc-BatCoV-512, TGEV, PRCV and FIPV. Genus Betacoronavirus; type species: Subgroup A: Murine coronavirus: HCoV-OC43, HCoV-HKU1, ECov, PHEV, BCoV, and MHV. Subgroup B: SARSr-CoV and SARSr-Rh-BatCoV HKU3. Subgroup C: Pi-BatCoV HKU5, Ty-BatCoV HKU5 and MERS-CoV. Subgroup D: Ro-BatCoV HKU9. IBV and TCoV in the genus Gammacoronavirus and MunCoV HKU13, BuCoV-HKU11, BuCoV-HKU17, ThCoV-HKU12, and PdCV in the genus Deltacoronavirus.
The mechanism of pathogenesis
The coronavirus, which is much smaller than the cell, has some of the genetic elements (RNA) used to transfer information. A protective membrane called the capsid protects this genetic information. The capsid is embedded in a coating with a flagellum or crown that plays an important role in the virus entering the cells. Once the virus reaches the appropriate cells, the virus crowns act just like a key and attach to its receptor, and enter the cell. The glycoprotein-s virus with two subunits S1 and S2 (mediating virus-cell membrane fusion), uses the same receptor, the same angiotensin-converting enzyme-2 (ACE-2), to enter the cell and is therefore not identified as an alien. In the normal human lung, ACE-2 is expressed in type 1 and 2 alveolar epithelial cells, with 83% of type-2 alveolar cells expressing ACE-2. ACE-2 are the host cell receptors that the S glycoprotein of severe acute respiratory syndrome coronavirus (SARS-CoV-2) binds to it. ACE-2 has been introduced into the human lower respiratory tract, the SARS-CoV-2 cell receptor. Men have higher ACE-2 levels in their alveolar cells than women(3). As previously mentioned, the prerequisite for a coronavirus attack on the host cell is to bind to the receptor. Upon receptor binding, the viral spike protein is broken down by acid-dependent proteolysis by cathepsin, TMPRRS, or furin proteinase and then integrates with the cell membrane. Compared to other coronavirus proteins, the spike protein has the most variable amino acid sequence, the strongest option among all coronavirus genes for adaptation to their hosts. Endocytosis is the mechanism of virus entry into the cell, the coronavirus genetic material (RNA) is released into the cell's cytoplasm, takes control of protein synthesis by taking over the ribosome, and encodes non-structural proteins. The Golgi apparatus completes its assembly by releasing the virus and releasing it by exocytosis. The main problem is that the production of particles in the endoplasmic reticulum network causes cell stress and death. The immune system begins to produce antibodies due to a backlog of external factors not present in the immune memory. At this point, the enzymes in the cell's nucleus find the antibody-producing code and send a copy, called messenger RNA (mRNA), to the ribosome. These results in body warming to make the virus harder to function, nosebleeds and sputum to drive dead cells, and bone weakness to produce white blood cells by the bone(4-6). Overexpression of ACE2 in various cells, such as lung AT2 cells, upper esophagus, epithelial cells, and ileum and colon absorptive enterocytes, may be involved in multinuclear infection of the new coronavirus 2019 (4, 5, 7). In a healthy person with a proper immune system, external agents are released after some time. Still, in complex and acute conditions, the immune cells responsible for triggering some chemical reactions may also cause lung damage. Combining these fluids with dead cells disrupts the flow of oxygen to the bloodstream and eventually leads to shortness of breath or suffocation. As well as focusing entirely on the body to cleanse the virus of some of the weakened organisms in the body, such as bacteria that were not previously grown, take advantage of the opportunity, which may disrupt some vital organs of the body such as the kidneys and can even lead to death (3, 8, 9). On the other hand, the stress of the endoplasmic reticulum and activation of the protein response can be surprisingly involved in the proliferation and pathogenesis of coronavirus during disease and infection (10).
Coronavirus and immune system
Virus infection is usually associated with upper respiratory tract inflammation, fever, headache, and cough. SARS-CoV-2 and MERS-CoV infection can be asymptomatic in the early stages but can later cause pneumonia, kidney failure, and even death. Pneumonia appears to be the most common manifestation of infection, characterized in the early stages by fever, cough, shortness of breath, and bilateral infiltration in chest imaging. In addition to respiratory symptoms, gastrointestinal symptoms (e.g., nausea and diarrhea) have been reported in some patients, but these are relatively rare. In patients with COVID-19, the number of white blood cells can vary. Leukopenia, leukocytosis, and lymphopenia have been reported, although the most common manifestation is lymphopenia. Pulmonary parenchymal abnormalities in patients have been observed, such as bilateral patchy shadows or ground-glass facades, as well as high levels of aminotransferase. At admission time, many pneumonia patients have normal serum procalcitonin levels. However, their level is likely higher in people needing intensive care (ICU). The failure of the immune system in response to coronaviruses can lead to immunopathology and death. Chemotactic factors (such as Interleukin8, C5a, and C3a) are essential for the immune response to the virus because these factors play a role in the recruitment of leukocytes to the lung, and any change in these factors can impair immune function. Immune defects or abnormalities can increase virus replication. Excessive immune system activity can also cause tissue damage (1).
Innate immune response
The immune system is incorporated via innate immune receptors: RIG-I-like receptors, C-type lectin-like receptors, Toll-like receptors, and NOD-like receptors can detect coronavirus. Type I interferons are the first line of defense against viruses. These viruses escape the host immune system using various methods, such as JAK-STAT signaling inhibition induced by type I interferon. One of the most significant features of enveloped coronaviruses is their ability to infect macrophages and activate type I interferons (IFN-α/β), which stimulate the immune system, such as natural killer cells and macrophages, and increase body resistance. Coronavirus infection has also been displayed to delay IFN signaling and lead to acute disease. Alpha and beta interferons are synthesized by many cells after coronavirus infection, preventing the spread of the virus, increasing the incidence of MHCi on uninfected cells, increasing their resistance to NK cells, and sensitizing them to CTL. On the other hand, the virus interferes with the differentiation and function of dendritic cells and thus affects specific immune responses (1). Pathogen-associated marker molecules (PAMPs) are responsible for detecting virus RNA in the innate immune system. TLR3, TLR7, and cytosolic RNA receptor receptors such as RIG-I and MDA5 activate NF-κB and IRF3 signaling cascades, which ultimately lead to increased expression of IFN-I in primary viral infections. The coronavirus can escape the immune system. Studies of the SARS and MERS viruses have shown that these viruses are capable of suppressing IFN-I production responses directly and indirectly. Ubiquitinase and reduction of STAT1 phosphorylation, which are involved in IFN-I signaling, are among the mechanisms of the virus suppressing innate immune responses. Those infected with coronavirus have higher levels of proinflammatory cytokines such as IP-10, MCP-1, MIP-1A, and TNFα, and the deterioration of patients is directly correlated with the levels of these cytokines.
Dedicated immune response
TCD4+ cells play an important role in immunity against the coronavirus by stimulating the production of specific coronavirus antibodies by B cells. Cytotoxic TCD8+ cells can kill infectious cells. IL-17 family cytokines are involved in infection control by inviting monocytes and neutrophils to the site of infection and activating the production of other inflammatory cytokines, for example, IL-1, IL-6, IL-8, and tumor necrosis factor α (TNF-α). Besides, MERS-CoV induces apoptosis in T cells. Lethal T cells directly attack the virus-infected cells. Then, a special protein called perforin is produced that creates pores in the infected cells and causes them to die, called cellular immunity (1).
Humoral immunity
Humoral immunity, as part of a dedicated defense, fights viruses in body fluids, in which B lymphocytes are involved. When B lymphocytes first bind to the antigen, they grow, divide, and transform into plasmocytes and B cells of memory. The virus uses its surface proteins as an adhesion factor to enter host cells through a specific receptor called dipeptidyl peptidase-4. This receptor is a key factor in activating the immune system in affected individuals(1). The target is focusing on more specific and specific arms, such as the controlled production of oxygen radicals or the production of cytokines such as IL-1, IL-4, IL-6, IL-8, and IL-21 which are more feasible (11). Finally, more studies need to be done to better understand the features of inflammatory responses associated with coronavirus infection and its relationship to the timing of treatment interventions. Xufang Denga and colleagues (2017) showed that modulating the stability and activity of coronavirus nsp15 protein in macrophages is essential for evading host dsRNA sensors and proposed a strategy for producing live attenuated vaccines (12).
The effect of coronavirus on apoptosis and autophagy
Apoptosis is an important defense mechanism for controlling virus infection.
Many viruses can actively cause apoptosis, which is a way that undeveloped and non-lytic viruses can kill host cells by limiting inflammatory and other immune responses(13). Apoptosis is operated either in the intrinsic or extrinsic pathway. The B cell lymphoma 2 family proteins are organized by the intrinsic pathway. Among B cell lymphoma two-family proteins, BAX and BCL2-antagonist/killer 1(BAK proteins) are pro-apoptotic, increasing the mitochondrial outer membrane permeability. Mcl-1, Bcl-xL, and Bcl2 proteins are Bcl2-like proteins as anti-apoptotic factors and prevent mitochondrial outer membrane permeability. As the mitochondrial outer membrane permeability increases, at first, cytochrome-c is released, then the apoptosome is formed, and eventually, it activates caspase 7/3. In the extrinsic pathway, the Connection of FasL and TNF-α as death ligands to the cellular surface death receptors creates a set of death-induced signals and then activates Caspase-8 (14). Some viruses and viral products can inhibit apoptosis, and others are involved in its induction. Parvovirus, Adenovirus, Papillomaviruses, Chicken Anemia Virus, Simian Virus 40, and human immunodeficiency virus type-1/Simian immunodeficiency virus are viruses that induce apoptosis. These viruses induce apoptosis by their specific proteins (13).
Adenovirus, African swine fever virus, Bovine papillomavirus, Baculovirus, Cowpox virus, Epstein-Barr virus, Hepatitis B virus, Human Cytomegalovirus, Hepatitis C virus, Herpes simplex virus, Gamma herpesviruses, Myxoma virus, Kaposi's sarcoma herpesvirus, Simian Virus 40, and Vaccinia virus are viruses that inhibit apoptosis. These viruses use p53 inactivation to prevent apoptosis (13). In line with other studies, TS Fung et al., in their study in 2019, revealed that simultaneously with virus replication, autophagy activity, apoptosis, and oxidative stress are well mediated in epithelial cells infected with gastroenteritis virus (15, 16). However, another study examining two cell types showed that infecting swine testicular cells with Transmissible Gastroenteritis Virus or infecting intestinal porcine enterocytes with isolated neonatal jejunum from a neonatal piglet with porcine epidemic diarrhea virus, together with autophagy activity, suppresses virus replication (15, 17, 18). On the other hand, the p53 tumor suppressor protein is required for signal propagation in apoptosis. In many cases, the p53 tumor suppressor protein mediates this cellular suicide, although the precise mechanism of its function has not yet been elucidated. Both p53-mediated transcriptional activation and a p53 function without transcriptional activation appear to be involved in apoptosis (13). Bcl-2 family members also promote apoptosis in many cases. Recent laboratory studies indicate that virus-induced apoptosis plays a critical role in cell death and killing cytotoxic T-lymphocytes infected with the virus (13, 19). Cytotoxicity is usually characterized by the activity of CD8+T cells and CD4+T cells (20). Cytotoxic T lymphocytes with two secretory and membrane pathway mechanisms (including the perforin-granzyme and Fas-FasL pathway) can play a role in toxicity and cell death. In the membrane pathway of perforin-granzyme, upon activation of the T-cell receptor, the target cells lyse and simultaneously release lytic granules that contain perforin and serine proteases (21). Whereas, in the second membrane pathway, the Fas/Fas Ligand pathway, the target cells become toxic when Fas Ligand expression is specifically observed in activated T cells and at the same time induces Fas binding to the target cells (21-24). Research has indicated that alpha and beta coronavirus infection can promote autophagy (25-27). Helena J. Maier, et al. in 2012 showed that protein 1Blight chain 3 (as part of the ERAD tuning pathway), suggesting induction of autophagy in coronavirus infection by the punctate virus (28).
Ying-Xim Tan et al. 2007 showed that Bcl-XL overexpression can occlusions apoptosis induced by 7a structural protein via interfering with the pro-survival Bcl-XL function in different stages of the infection cycle (29). Structural protein 7a is one of the severe acute respiratory syndrome coronavirus proteins and plays a significant role in some functions such as coronavirus pathogenesis, prompts apoptosis, cell cycle arrests, and stimulates the production of pro-inflammatory cytokines (29) (Figure 4).

Figure 4. Infection with Human Corona Virus induces apoptosis and its modulating mechanisms. The apoptosis pathway contains the stimulation of death receptors on the cell surface, followed by a caspase cascade. Apoptosis comprises two intrinsic pathways and an extrinsic pathway. The intrinsic apoptosis cascade involves the proliferation of pro-apoptotic mediators, triggering both caspases 9 and 3. The extrinsic cell death pathway is facilitated by death receptors. Fas ligand binding activates the death receptor. The Bcl-2 family proteins are organized by the intrinsic pathway. Both BAX and BAK proteins are pro-apoptotic. Also, Bcl-xL and Bcl2 proteins are anti-apoptotic factors and can prevent mitochondrial outer membrane penetrance.
Autophagy is a major intracellular pathway for destroying and recycling long-lived proteins and damaged organelles that occur through various pathways, including macroautophagy, macrophage, and autophagy mediated by chaperones. Also, Autophagy is involved in many physiological processes such as cell growth, aging, cell death, stress response, and immune response in cells infected with viral infection (30). Activation of protein responses in the cell can publicly control and regulate a wide range of cellular signaling pathways like mitogen-activated protein kinase actuation, apoptosis, autophagy, innate immune response, and so on. Beclin-1 and LC3 genes play a central role in autophagy among autophagy-regulating genes. Beclin-1 is involved in signaling pathways and the initiation phase of auto-phagosome formation, where interaction with PI3PK and hvp34 is essential. Beclin-1 cooperates directly with the Bcl-2 protein (31). The proliferation of the Middle East Respiratory Syndrome can block the fusion of autophagosomes and lysosomes by decreasing Beclin-1 levels (32). By inhibiting SKP2, Beclin-1 depletion is observed and can also decrease Beclin-1 degradation and increase autophagic flux. Eventually, Nils C. Gassen et al. (2019) suggested that the S-phase kinase-associated protein could be used as an engaged goal for drug production against coronavirus. They utilized conditions sensitive to autophagy (32). Coronavirus envelope (E) protein is a very small and integral membrane protein that plays an important role in various aspects of the viral replication cycle: including assembly, induction of membrane curvature to scission or budding, envelope formation, pathogenesis and release to apoptosis, inflammation and even self-restraint or autophagy. In 2019, David Schumann et al. suggested that, according to extensive research on COVID-19, available data indicate that envelope (E) protein of CoVs plays a key role in the viral life cycle, and the CoVs lacking E can be hopeful vaccines candidates for both human CoVs and enzootic CoVs (33). Lu et al. 2020 have confirmed that in addition to antiviral and antibiotic interactions, photoreceptors, RNA synthesis inhibitors, and traditional Chinese medicine can possibly be useful in treating Covid-19 (34). Yu F et al., 2020, stated that a peptide based on angiotensin-converting enzyme-2, a 3CLpro-1 inhibitor, and a new vinyl sulfone proteinase inhibitor, in theory, appears to have the potential of antiviral activity against novel coronavirus-2019 (35). W Liu et al.2020 stated that four important enzymes are essential for the pathogenesis of the virus: First, the rod protein that facilitates virus entry through the Angiotensin-converting enzyme 2 receptors to the host cell surface receptor. Second, Coronavirus Main Proteinase, third, Papain-like protease involved in the assembly of new viruses; and Finally, RNA Dependent RNA polymerase facilitates genomic replication of the coronavirus RNA. nCoV-2019, SARS RdRp, and CLpro are very similar in sequence, making it possible to use small drug molecules previously based on SARS-CoV proteins, such as Remdesivir and 3LCpro-1, make it possible to treat the new coronavirus (36). Many data suggest that the vaccine should incorporate components of the virus's structural proteins that elicit an effective immune response. Therefore, in the design of the vaccine, it seems that the production of a monoclonal antibody against this protein can provide helpful immunity. Targeted immunotherapy can also be one of the therapeutic strategies. In this regard, glucocorticoids have been widely used to treat SARS to control lung infection and inflammation. Except for the pathogenic stage of the disease, the body's inflammatory response plays a key role in causing SARS-related lung injury. In the absence of effective treatments, the best way to deal with the COVID-19 Epidemic is to control the sources of infection. Strategies include early detection, reporting, isolation, quarantine, and supportive treatments. Timely dissemination of epidemic information and maintenance of social order are also effective. At the individual level, protective measures, including personal hygiene, medical mask wear, adequate rest, and proper room ventilation, can effectively prevent modern coronavirus-2019 infection (37).
The vaccine against the 2019 novel coronavirus SARS-CoV-2
Currently, many researchers are working to develop it according to the virus's known molecular pathways. It is known that both SARS-CoV and SARS-CoV-2 use the ACE-2 receptor to enter human cells, which must be considered in attempts to produce the vaccine to treat or stop the virus (38). Lately, some companies have claimed the product of the vaccine, including Pfizer–BioNTech, and Moderna. However, there is no information about the true efficacy of these vaccines in the population of different countries, and further attempts are needed to end the pandemic.
Several ideas can help make a vaccine against the virus
1- Complete the virus using a dead virus or an inactivated SARS-CoV-2 virus that can provide a very fast immune response in a living organism (39). Vaccines made Complete the virus have innate immunity and the ability to stimulate toll-like receptors, including toll-like receptors 3, toll-like receptors7/8, and toll-like receptors9 (40). However, such vaccines require many experiments to be used in the human phase to confirm safety.
2- Making the vaccine using a special subunit of the virus that is sensitive to it. However, for the SARS-CoV-2 virus, research on S-spike protein as a subunit that givpid nanoparticle dispersion containing messenger RNA (44, 45)
7- Vaccination using a lentiviral vector, pathogen-specific artificial antigen-presenting dendritic cells (46)
8- Vaccination using lentiviral minies the virus the ability to bind to ACE2 enzyme receptors and inhibit virus function (38).
3- One of the newest ways to make a vaccine is to use DNA or RNA(41).
4- Preparing the vaccine using the recombinant adenovirus type 5 vector method (42)
5- Preparing the vaccine using an adenovirus vector (43)
6- Preparing the vaccine using li
gene vaccine, dendritic cells modified with a lentiviral vector (43)
9- Preparing the vaccine using DNA plasmid delivered by electroporation (47, 48).
Such vaccines can be tested experimentally using these nine methods for safety and efficacy in animals and in the next phase on humans as volunteers.
Conclusions
The prevalence of COVID-19 has become a worldwide clinical threat. Even though much progress has been made in diagnosing COVID-19, there is still a great deal to be discovered about this virus regarding its function and inhibition. By further investigating the function of proteins and envelope of coronavirus, effective antiviral agents and vaccination against the virus should be developed. These studies will help to further decrease mortality and transmission rates in patients infected with the new coronavirus-2019. This article is an overview of the collection of information and the identification of coronavirus functional mechanisms for developing and producing vaccines against it.
Authors’ contributions
AM wrote the original draft, DT mainly designed and supervised the project, LOR reviewed and edit the manuscript. All authors reviewed and approved the final version of the manuscript.
Acknowledgments
Special thanks to the Department of Urology, UroScience, State University of Campinas, Unicamp and Pontifical Catholic University of Campinas, PUC-Campinas, Campinas, SP, Brazil
Conflict of interest
The authors declare that there is no conflict of interest.
Funding
The authors received no financial support for this research.
Ethical Statements
Not applicable
Data availability
Data will be provided by the corresponding author on request.
Abbreviations
ACE-2 Angiotensin-converting enzyme-2
APAF1 Apoptotic peptidase-activating factor 1
BAD Bcl2-associated agonist of cell death
BAX Bcl2-associated X
Bcl2 B cell lymphoma 2
FasL Fas ligand
IFN-α/β Interferons α/β
IL-1 Interleukin 1
MCP-1 Monocyte chemoattractant protein-1
MDA5 Melanoma differentiation-associated protein 5
MERS Middle Eastern Respiratory Syndrome
NF-κB Nuclear factor-kappa B
PdCV Porcine deltacoronavirus
PI3PK Phosphoinositide 3-kinases
PUMA P53-upregulated modulator of apoptosis
RIG-I Retinoic acid-inducible gene I
SARS Severe Acute Respiratory Syndrome
TLR Toll-like receptor
TNF-α Tumor necrosis factor-alpha